The goal of NetCoupler is to estimate potential causal links between a set of -omic (e.g. metabolomics, lipidomics) or other high-dimensional metabolic data as a conditional dependency network and either a disease outcome, an exposure, or both. These potential causal links are classified as direct, ambigious, or no effects. This algorithm is largely meant to be used with -omic style data to generate the networks and while theoretically non-omic data could be used, we have not tested it in that context. Given the algorithms nature, it’s primarily designed to be used for exploration of potential mechanisms and used to complement other analyses for a research question. It could also be used to confirm a pre-specified and explicit hypothesis, similar to how structural equation models are used. However, this might be a more niche use.
Why or when might you want to use NetCoupler?
- You are interested in asking a research question on how some factor might influence another factor and how it might mediate through a metabolic network.
- If you want to explore how a factor might influence a metabolic network or how a metabolic network might influence a factor.
- You have an -omic dataset and want another method to explore how it relates to your variable of interest.
Basically, if you’re research question or objective has the general form of:
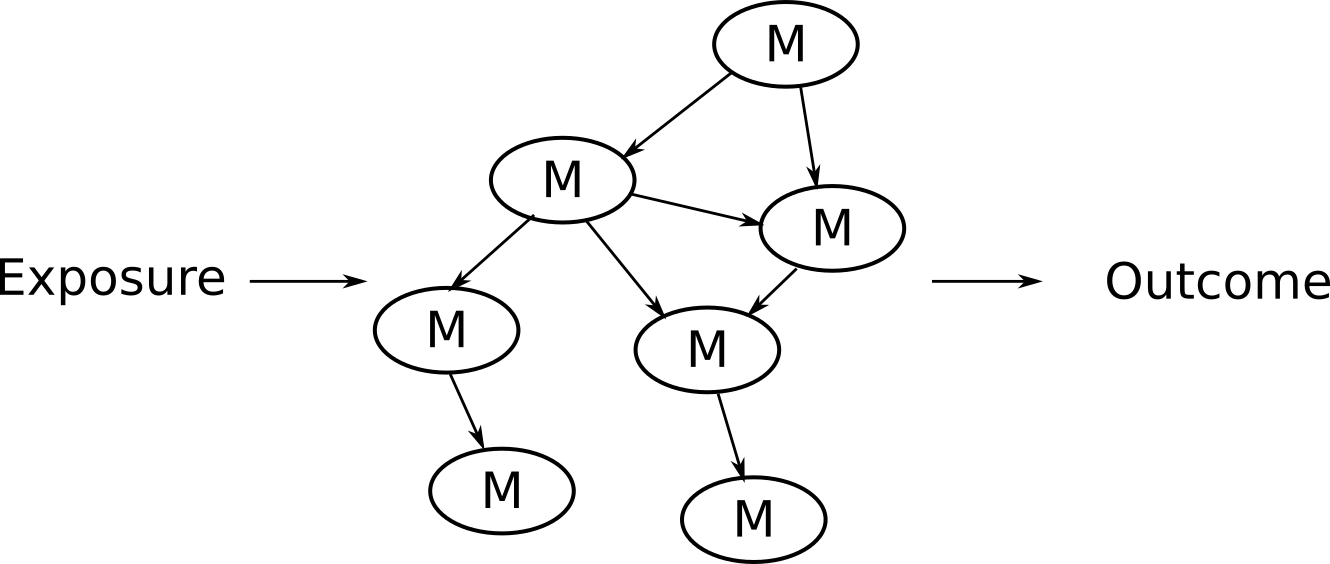
… So that you can ultimately have an answer that looks like:
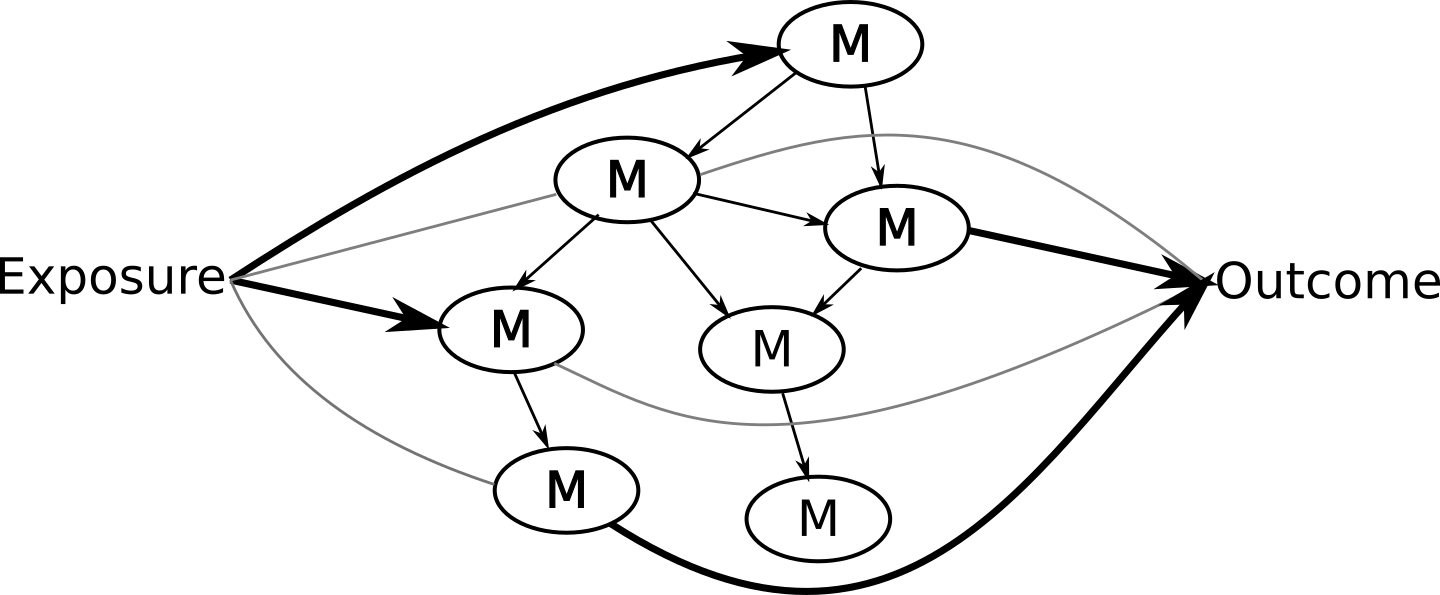
There are a few vignettes available in this package:
-
Get Started (
vignette("NetCoupler")) describes a simple overview of how and when to use NetCoupler, as well as a basic explanation of some of the components of NetCoupler. -
Examples with different models (
vignette("examples")) lists different models we’ve tested that work with NetCoupler. If you have tried a model out that isn’t listed and seen success, let us know by opening an Issue or submitting a Pull Request (see the contributing guidelines for instructions on doing this).
Installation
To install the official CRAN version, use:
install.packages("NetCoupler")To install the development version, use:
# install.packages("remotes")
remotes::install_github("NetCoupler/NetCoupler")Contributing and Code of Conduct
Checkout the guidelines for details on contributing. Please note that the ‘NetCoupler’ project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.