Examples of using NetCoupler with different models
Source:vignettes/articles/examples.Rmd
examples.RmdThis article contains a list of models that you could use in NetCoupler as well as some published real-world examples of it being used.
There are some caveats to most of these examples, except for when a section explicitly indicates otherwise:
- Most of the models do not include further adjustment for confounders.
- Most models only show the outcome-side estimation, for brevity. However, the code for running exposure-side estimation is almost the exact same.
Across the different models, there are some general features, which is described in more depth in the Getting Started article.
Pre-processing
First load up the packages.
-
Pre-processing by standardizing variables, since large differences in the values of variables can impact on the results of the algorithm.
standardized_data <- simulated_data %>% nc_standardize(starts_with("metabolite")) -
Estimating the network structure to identify links between the metabolic variables. This network needs to be converted into an edge table that has two columns, one for the
source_nodeand another for thetarget_node.metabolite_network <- simulated_data %>% nc_standardize(starts_with("metabolite")) %>% nc_estimate_network(starts_with("metabolite")) edge_table <- as_edge_tbl(metabolite_network) edge_table -
If adjusting for confounders in the main models, these also need to be included when estimating the network. To do this, the metabolic data needs to be regressed on to the variables that will be standardized.
metabolite_network <- simulated_data %>% nc_standardize(starts_with("metabolite"), regressed_on = "age") %>% nc_estimate_network(starts_with("metabolite"))
Outcome vs exposure side
Let’s revisit this image:
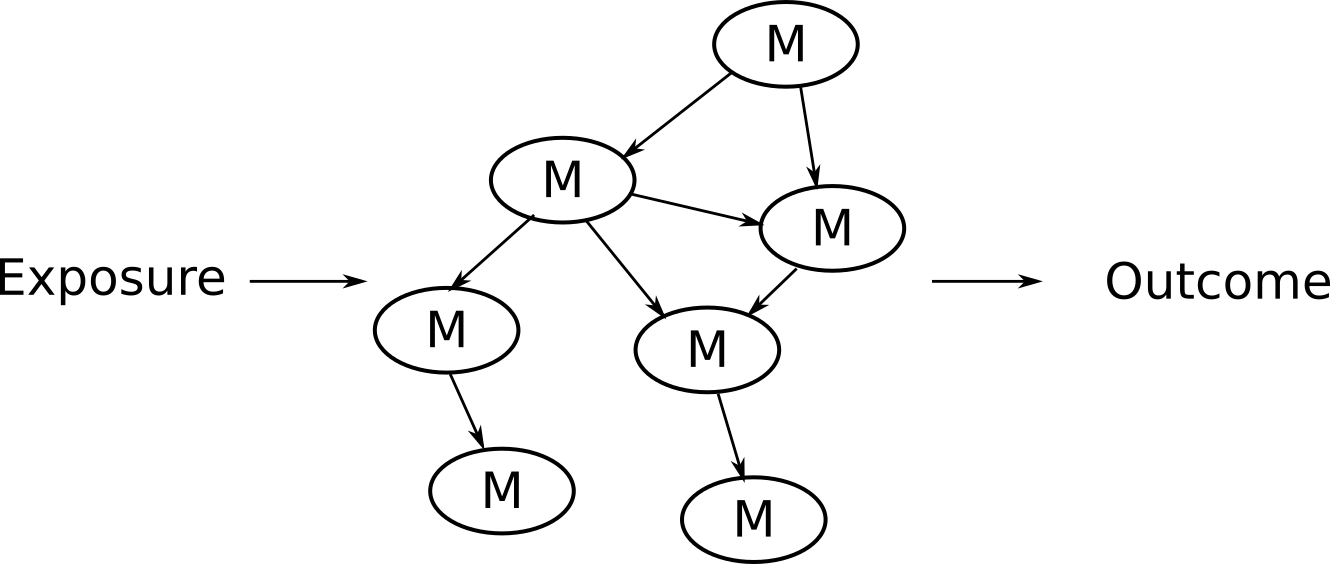
All types of models can be used for either the left hand side of this
graph (the exposure side) or the right hand side (the outcome). If we
want to estimate the links on the exposure side, where we’re interested
in how a variable might influence the network, we would use the
nc_estimate_exposure_links() function.
standardized_data %>%
nc_estimate_exposure_links(
edge_tbl = edge_table,
exposure = "exposure",
model_function = lm
)If we are interested the outcome side, where we want to know how the
network might influence the outcome, we would use the
nc_estimate_outcome_links() function.
standardized_data %>%
nc_estimate_outcome_links(
edge_tbl = edge_table,
outcome = "outcome_continuous",
model_function = lm
)Linear regression
This is the easiest and will probably be the most commonly used
modeling method used when running NetCoupler. Adding additional
arguments to settings to the lm() (or glm())
function can be done by using the model_args_list
argument.
lm_results <- standardized_data %>%
nc_estimate_outcome_links(
edge_tbl = edge_table,
outcome = "outcome_continuous",
model_function = lm
)Logistic regression
Binary Logistic Regression
Probably the second most common model would be the binary classic
logistic regression. Unlike the linear regression modeling above, we
need to use the model_arg_list argument in order to tell
glm() to use the binomial method for model estimation.
glm_bin_results <- standardized_data %>%
nc_estimate_outcome_links(
edge_tbl = edge_table,
outcome = "outcome_binary",
model_function = glm,
model_arg_list = list(family = binomial),
exponentiate = TRUE
)Cox proportional hazards regression
With Cox models, the response/y variable usually needs to be a
survival::Surv() object. While you can use this function in
the outcome/exposure argument of the
nc_estimate_outcome_links() or
nc_estimate_exposure_links() functions, to keep the code
and output a bit cleaner, we recommend creating the survival object
beforehand with mutate().
library(survival)
cox_surv_data <- standardized_data %>%
mutate(surv_object = Surv(
time = age,
time2 = age + outcome_event_time,
event = outcome_binary
))
coxph_results <- cox_surv_data %>%
nc_estimate_outcome_links(
edge_tbl = edge_table,
outcome = "surv_object",
# Can also use Surv directly.
# outcome = "Surv(time = time_start, time2 = time_end, event = outcome_binary)",
model_function = survival::coxph
)You might want to add a clustering to calculate robust standard errors or to add a strata variable. You add these directly to the adjustment variable argument.
coxph_results_cluster <- cox_surv_data %>%
mutate(age = as.integer(age)) %>%
nc_estimate_outcome_links(
edge_tbl = edge_table,
outcome = "surv_object",
adjustment_vars = c("strata(age)", "cluster(id)"),
model_function = survival::coxph
)